People
Filters
Projects: LiSyM Pillar I: Early Metabolic Injury (LiSyM-EMI), LiSyM PALs, LiSyM network, LiSyM-Krebs Partnering, Forschungsnetzwerk LiSyM-Krebs, DEEP-HCC network
Institutions: Zentrum für Informationsdienste und Hochleistungsrechnen (ZIH), Technische Universität Dresden
 https://orcid.org/0000-0003-0137-5106
https://orcid.org/0000-0003-0137-5106
Projects: LiSyM Core Infrastructure and Management (LiSyM-PD), LiSyM Pillar I: Early Metabolic Injury (LiSyM-EMI), LiSyM Pillar II: Chronic Liver Disease Progression (LiSyM-DP), LiSyM Pillar III: Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF), LiSyM Pillar IV: Liver Function Diagnostics (LiSyM-LiFuDi), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), LiSyM PALs, LiSyM network, LiSyM-Krebs Partnering
Institutions: HITS gGmbH
 https://orcid.org/0000-0002-8683-7084
https://orcid.org/0000-0002-8683-7084
Data management and standardization expert for systems biology and systems medicine, responsible for the data management user requirements and user contacts within the German LiSyM network (Liver Systems Medicine: http://lisym.org/) and associated to the FAIRDOM team. Involved in different standardization initiatives and committees, i.e. COMBINE (http://co.mbine.org), ISO/TC 276 Biotechnology (https://www.iso.org/committee/4514241.html), European COST action CHARME (http://www.cost-charme.eu) and ...
Projects: LiSyM Pillar II: Chronic Liver Disease Progression (LiSyM-DP), LiSyM Pillar III: Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF), LiSyM network, LiSyM PALs, LiSyM-Krebs Partnering, C-TIP-HCC network, Forschungsnetzwerk LiSyM-Krebs
Institutions: Molecular Hepatology - Dept of Medicine II - Medical Faculty Mannheim - Heidelberg University
Projects: LiSyM Core Infrastructure and Management (LiSyM-PD), LiSyM network, LiSyM PALs, LiSyM Pillar II: Chronic Liver Disease Progression (LiSyM-DP), LiSyM Pillar I: Early Metabolic Injury (LiSyM-EMI), LiSyM Pillar IV: Liver Function Diagnostics (LiSyM-LiFuDi), LiSyM Pillar III: Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF)
Institutions: HITS gGmbH
Projects: LiSyM PALs, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), LiSyM Pillar IV: Liver Function Diagnostics (LiSyM-LiFuDi), LiSyM network, LiSyM-Krebs Partnering
Institutions: Humboldt-Universität zu Berlin - Institute for Theoretical Biology (ITB)
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
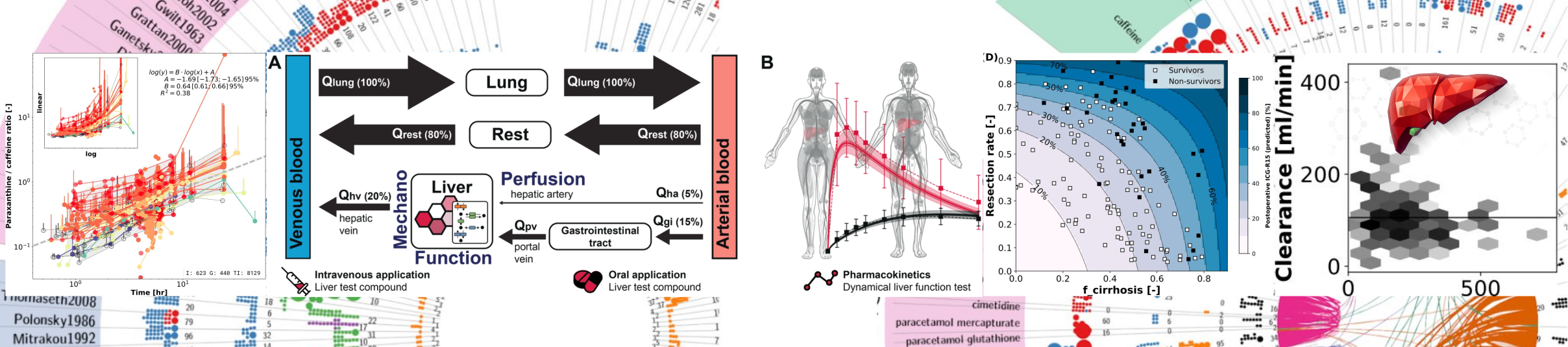
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: LiSyM PALs, LiSyM network, LiSyM Pillar I: Early Metabolic Injury (LiSyM-EMI), LiSyM Pillar II: Chronic Liver Disease Progression (LiSyM-DP), LiSyM Pillar IV: Liver Function Diagnostics (LiSyM-LiFuDi), LiSyM Core Infrastructure and Management (LiSyM-PD), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), The Hedgehog Signalling Pathway (LiSyM-JGMMS), DEEP-HCC network, C-TIP-HCC network, Default_Project
Institutions: HITS gGmbH
 https://orcid.org/0000-0003-3540-0402
https://orcid.org/0000-0003-3540-0402
I am a researcher at the Scientific Databases and Visualization Group at Heidelberg Institute for Theoretical Studies (HITS) , one of the developers of SabioRK - System for the Analysis of Biochemical Pathways - Reaction Kinetics (http://sabiork.h-its.org/) . I am working on design and maintenance of the information systems to store, query and analyse systems biology data; definition and implementation of methods for the integration of data from multiple sources. In SySMO-DB project ...
Projects: LiSyM PALs, LiSyM network, LiSyM Core Infrastructure and Management (LiSyM-PD)
Institutions: HITS gGmbH
Projects: The Hedgehog Signalling Pathway (LiSyM-JGMMS), LiSyM PALs, LiSyM network, LiSyM Scientific Leadership Team (LiSyM-LT), LiSyM-Krebs Partnering, SMART-NAFLD, DEEP-HCC network, Forschungsnetzwerk LiSyM-Krebs
Institutions: Universität Leipzig - Medizinische Fakultät - Institut für Biochemie, Universität Leipzig - Sächsischer Inkubator für Klinische Translation (SIKT)
Group leader of the junior group: The Hedgehog Signalling Pathway (LiSyM-JGMMS)
Projects: LiSyM Core Infrastructure and Management (LiSyM-PD), LiSyM PALs, LiSyM network, LiSyM Scientific Leadership Team (LiSyM-LT), LiSyM-Krebs Partnering, Forschungsnetzwerk LiSyM-Krebs, C-TIP-HCC network, SMART-NAFLD, DEEP-HCC network, Default_Project
Institutions: HITS gGmbH
 https://orcid.org/0000-0002-4980-3512
https://orcid.org/0000-0002-4980-3512
I am a trained computer scientist (Habilitation) and physicist (Diploma). The topic of my Habilitation was distributed information retrieval of image data, i.e. how to find complex data via fuzzy queries. One key part of my work now is rendering data shared and FAIR (Findable, Accessible, Interoperable, Reusable) in order to make them useful for search and data integration. Since beginning of the Virtual Liver Network, through LiSyM I+II and now LiSyM-Cancer I am coordinator of the data management ...